Two-colour microarray experiments
At the moment Annotare only supports one type of two-colour workflow (see graphic below), where two samples are connected with one common raw data file, which includes both channels. If you select two-colour for your Annotare submission, it will expect that you provide one file for the two samples that were hybridised together.
The supported workflow has a 2:2:1:1 relationship between samples, labelled extracts, hybridisation assays and data files:
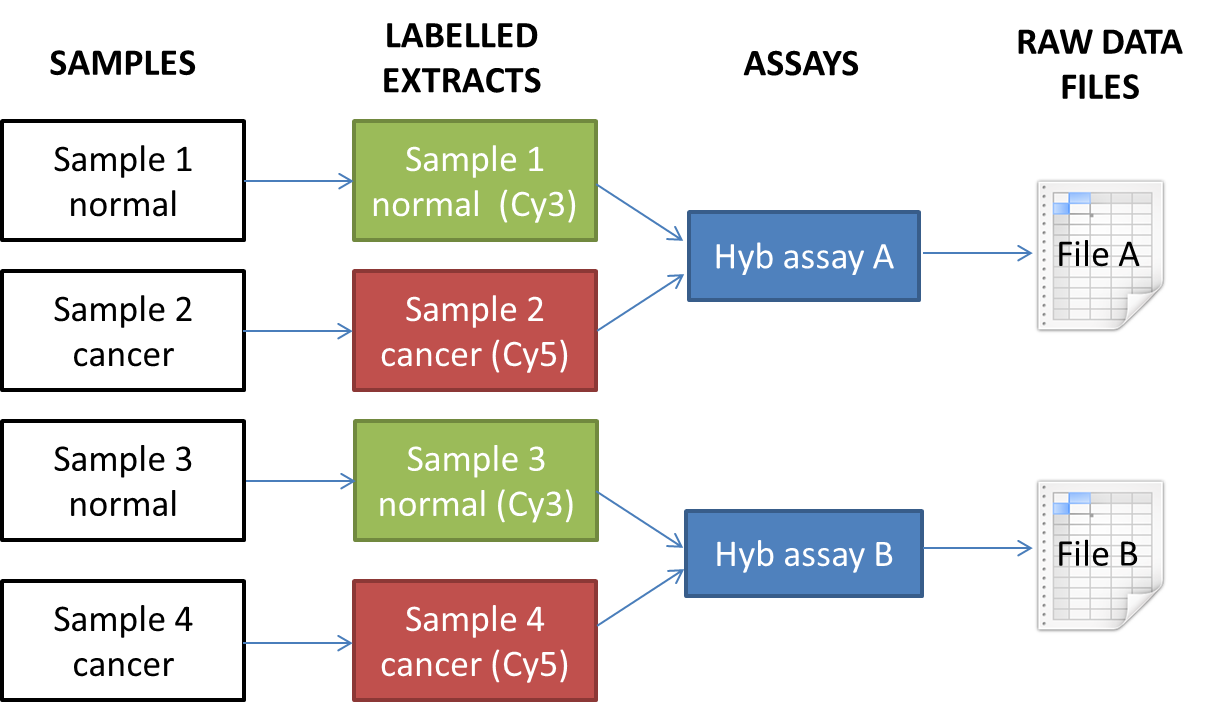
For this workflow, you need to tell us which label (fluorescent dye) is assigned to each extract, e.g. Cy3 or Cy5. By default Annotare uses your sample names as your extract names.
The experiment above would be represented like this through the different pages of Annotare forms:
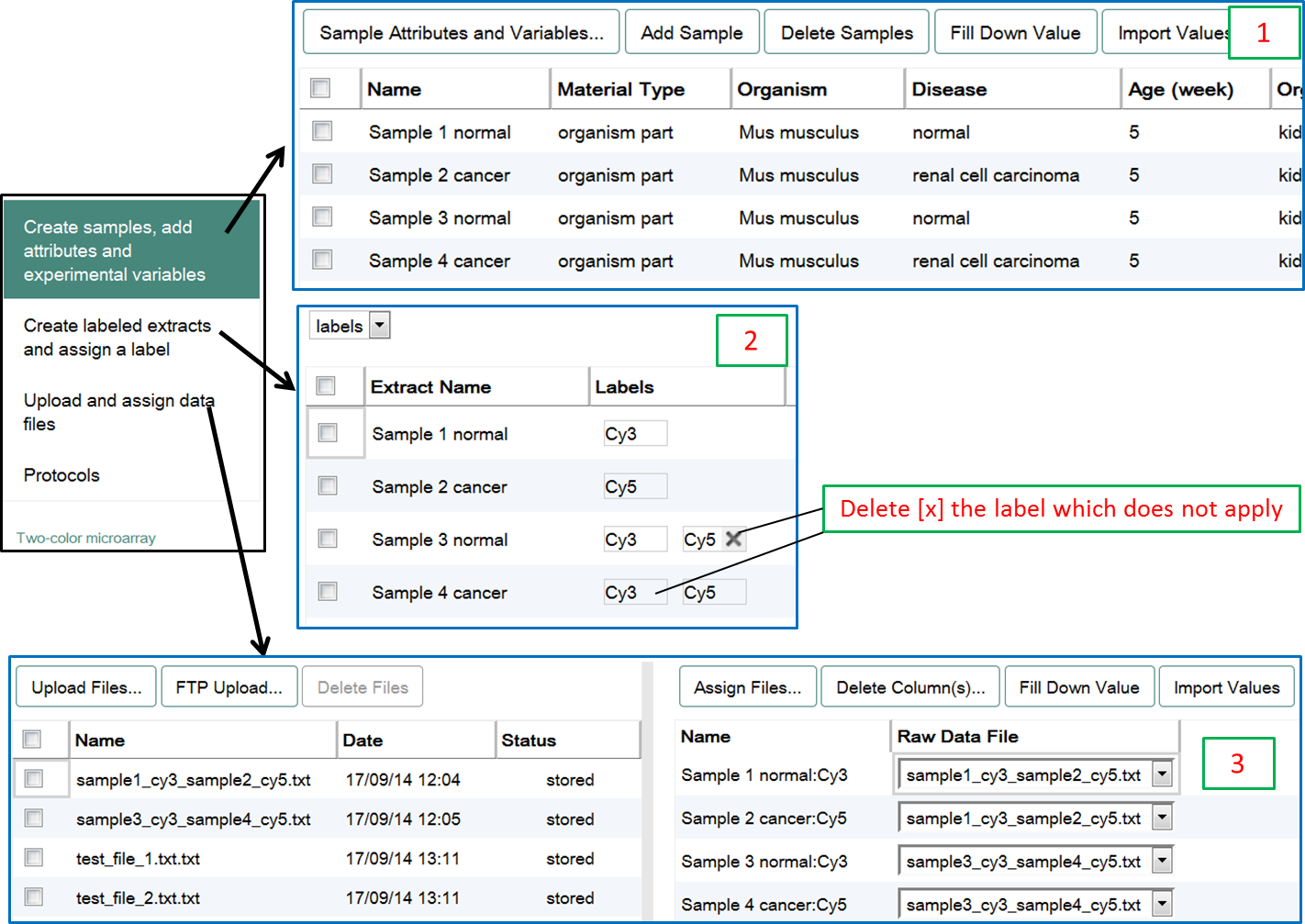
For more help on how to assign files for two-colour microarray experiments, see two-colour section of the Assign Files page.
Some two-colour microarray technologies generate two separate raw data files (usually one for each channel), which will cause Annotare validation to fail, if you connect a single file per sample. For such technologies follow the guidance below:
- If you have separate files for each channel, e.g. idat files from Illumina BeadChips, please select the "methylation microarray" template for your Annotare submission. You should not receive any file assignment error within this template. Make sure to select the correct experiment type (e.g. "genotyping by array" or "transcription profiling by array"), complete the Annotare form normally, and connect the files and samples as it makes sense for your experiment.
- If your two-colour experimental design features a common reference sample that is used in more than one hybridisation, please see this file assignment help for two-colour experiments for the workaround that needs to be applied in this case.